Researchers uncover that a spot of molecular glue and a well timed twist assist a bacterial enzyme convert carbon dioxide into carbon compounds 20 occasions sooner than plant enzymes do throughout photosynthesis. The outcomes stand to speed up progress towards changing carbon dioxide into a wide range of merchandise.
Carbon fixation, or the conversion of carbon dioxide from the air into carbon-rich biomolecules, is important for vegetation’ survival. That’s the entire level of photosynthesis, and a cornerstone of the huge interlocking system that cycles carbon by means of vegetation, animals, microbes, and the ambiance to maintain life on Earth.
The carbon-fixing champs, nevertheless, are soil micro organism, not vegetation. Scientists might be able to develop synthetic photosynthesis to transform greenhouse gasoline into fuels, fertilizers, antibiotics, and different merchandise if they will work out how sure bacterial enzymes perform an important step in carbon fixation 20 occasions faster than plant enzymes.
Now a crew of researchers from the Division of Vitality’sSLAC Nationwide Accelerator Laboratory, Stanford College, Max Planck Institute for Terrestrial Microbiology in Germany, DOE’s Joint Genome Institute (JGI) and the College of Concepción in Chile has found how a bacterial enzyme – a molecular machine that facilitates chemical reactions – revs as much as carry out this feat.
Quite than grabbing carbon dioxide molecules and attaching them to biomolecules one after the other, they discovered that this enzyme consists of pairs of molecules that work in sync, just like the arms of a juggler who concurrently tosses and catches balls to get the job achieved sooner. One member of every enzyme pair opens vast to catch a set of response substances whereas the opposite closes over its captured substances and carries out the carbon-fixing response; then, they swap roles in a continuous cycle.
A single spot of molecular “glue” holds every pair of enzymatic arms collectively to allow them to alternate opening and shutting in a coordinated method, the crew found, whereas a twisting movement helps hustle substances and completed merchandise out and in of the pockets the place the reactions happen. When each glue and twist are current, the carbon-fixing response goes 100 occasions sooner than with out them.
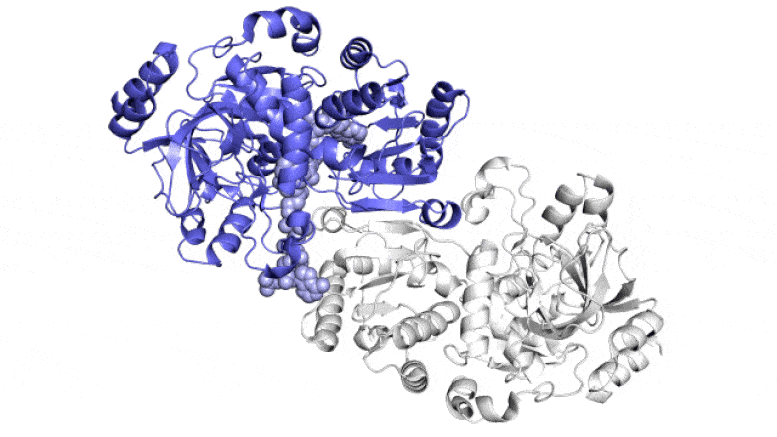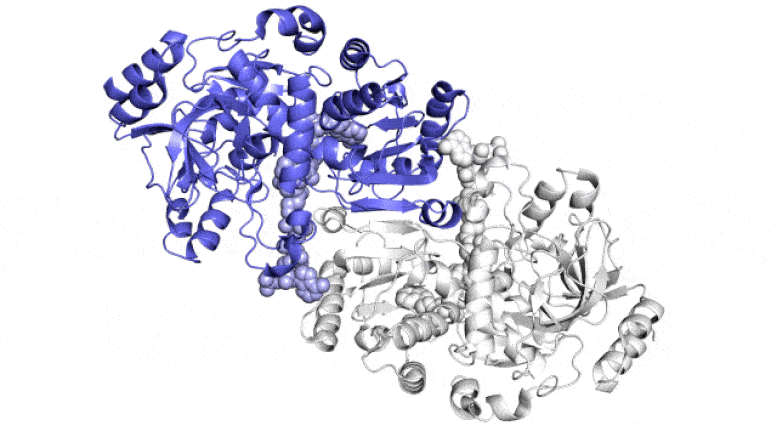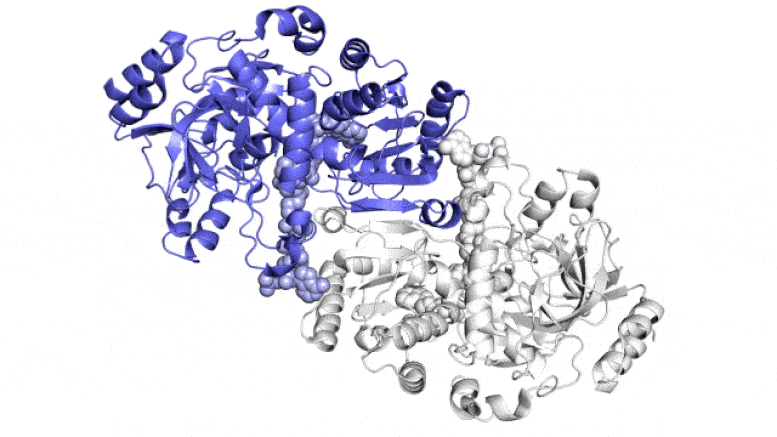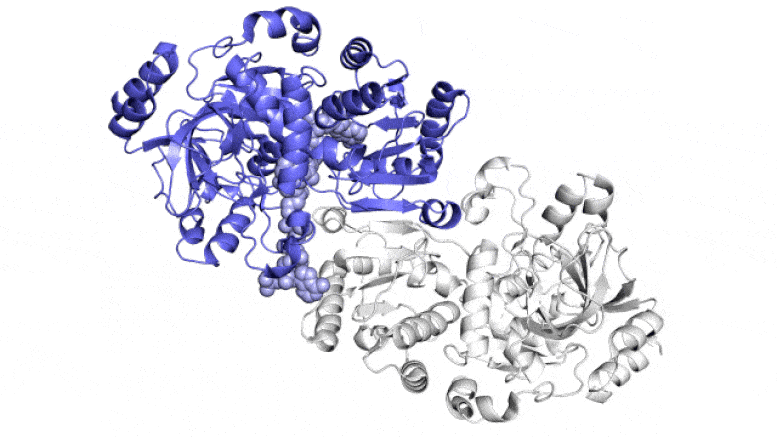
This animation exhibits two of the paired molecules (blue and white) throughout the ECR enzyme, which fixes carbon in soil microbes, in motion. They work collectively, just like the arms of a juggler who concurrently tosses and catches balls, to get the job achieved sooner. One member of every enzyme pair opens vast to catch a set of response substances (proven coming in from high and backside) whereas the opposite closes over its captured substances and carries out the carbon-fixing response; then, they swap roles in a continuous cycle. Scientists are attempting to harness and enhance these reactions for synthetic photosynthesis to make a wide range of merchandise. Credit score: H. DeMirci et al., ACS Central Science, 2022
“This bacterial enzyme is probably the most environment friendly carbon fixer that we all know of, and we got here up with a neat clarification of what it could possibly do,” stated Soichi Wakatsuki, a professor at SLAC and Stanford and one of many senior leaders of the research, which was revealed in ACS Central Science this week.
“A few of the enzymes on this household act slowly however in a really particular method to produce only one product,” he stated. “Others are a lot sooner and might craft chemical constructing blocks for all types of merchandise. Now that we all know the mechanism, we are able to engineer enzymes that mix the very best options of each approaches and do a really quick job with all types of beginning supplies.”
Enhancing on nature
The enzyme the crew studied is a part of a household known as enoyl-CoA carboxylases/reductases, or ECRs. It comes from soil micro organism known as Kitasatospora setae, which along with their carbon-fixing expertise may produce antibiotics.
Wakatsuki heard about this enzyme household half a dozen years in the past from Tobias Erb of the Max Planck Institute for Terrestrial Microbiology in Germany and Yasuo Yoshikuni of JGI. Erb’s analysis crew had been working to develop bioreactors for synthetic photosynthesis to transform carbon dioxide (CO2) from the ambiance into all types of merchandise.
As vital as photosynthesis is to life on Earth, Erb stated, it isn’t very environment friendly. Like all issues formed by evolution over the eons, it’s solely pretty much as good because it must be, the results of slowly constructing on earlier developments however by no means inventing one thing totally new from scratch.
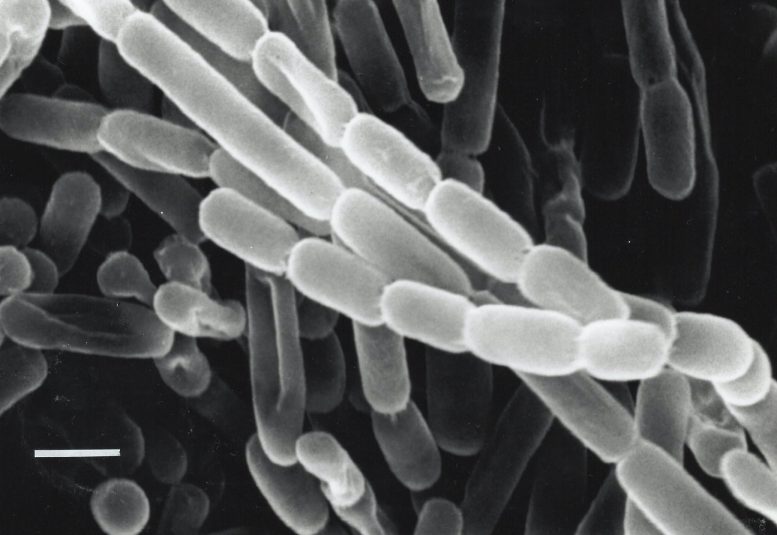
An in depth-up have a look at Kitasatospora setae, a bacterium remoted from soil in Japan. These micro organism repair carbon – flip carbon dioxide from their surroundings into biomolecules they should survive – because of enzymes known as ECRs. Researchers are searching for methods to harness and enhance ECRs for synthetic photosynthesis to provide fuels, antibiotics and different merchandise. Credit score: Y. Takahashi & Y. Iwai, atlas.actino.jp
What’s extra, he stated, the step in pure photosynthesis that fixes CO2 from the air, which depends on an enzyme known as Rubisco, is a bottleneck that bogs the entire chain of photosynthetic reactions down. So utilizing speedy ECR enzymes to hold out this step, and engineering them to go even sooner, may convey a giant increase in effectivity.
“We aren’t making an attempt to make a carbon copy of photosynthesis,” Erb defined. “We wish to design a course of that’s rather more environment friendly through the use of our understanding of engineering to rebuild the ideas of nature. This ‘photosynthesis 2.0’ may happen in residing or artificial techniques reminiscent of synthetic chloroplasts – droplets of water suspended in oil.”
Portraits of an enzyme
Wakatsuki and his group had been investigating a associated system, nitrogen fixation, which converts nitrogen gasoline from the ambiance into compounds that residing issues want. Intrigued by the query of why ECR enzymes had been so quick, he began collaborating with Erb’s group to seek out solutions.
Hasan DeMirci, a analysis affiliate in Wakatsuki’s group who's now an assistant professor at Koc College and investigator with the Stanford PULSE Institute, led the trouble at SLAC with assist from half a dozen SLAC summer season interns he supervised. “We practice six or seven of them yearly, and so they had been fearless,” he stated. “They got here with open minds, able to be taught, and so they did superb issues.”
The SLAC crew made samples of the ECR enzyme and crystallized them for examination with X-rays on the Superior Photon Supply at DOE’s Argonne Nationwide Laboratory. The X-rays revealed the molecular construction of the enzyme – the association of its atomic scaffolding – each by itself and when connected to a small helper molecule that facilitates its work.
Additional X-ray research at SLAC’s Stanford Synchrotron Radiation Lightsource (SSRL) confirmed how the enzyme’s construction shifted when it connected to a substrate, a form of molecular workbench that assembles substances for the carbon fixing response and spurs the response alongside.
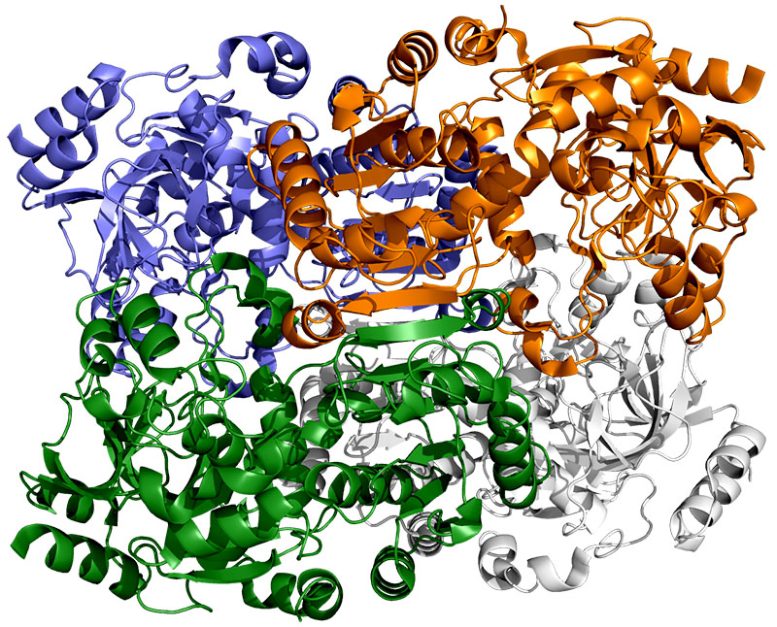
This depiction of ECR, an enzyme present in soil micro organism, exhibits every of its 4 similar molecules in a unique shade. These molecules work collectively in pairs – blue with white and inexperienced with orange – to show carbon dioxide from the microbe’s surroundings into biomolecules it must survive. A brand new research exhibits that a spot of molecular glue and a well timed swing and twist enable these pairs to sync their motions and repair carbon 20 occasions sooner than plant enzymes do throughout photosynthesis. Credit score: H. DeMirci et al., ACS Central Science, 2022
Lastly, a crew of researchers from SLAC’s Linac Coherent Gentle Supply (LCLS) carried out extra detailed research of the enzyme and its substrate at Japan’s SACLA X-ray free-electron laser. The selection of an X-ray laser was vital as a result of it allowed them to check the enzyme’s habits at room temperature – nearer to its pure surroundings – with nearly no radiation injury.
In the meantime, Erb’s group in Germany and Affiliate Professor Esteban Vo¨hringer-Martinez’s group on the College of Concepción in Chile carried out detailed biochemical research and intensive dynamic simulations to make sense of the structural information collected by Wakatsuki and his crew.
The simulations revealed that the opening and shutting of the enzyme’s two components don’t simply contain molecular glue, but in addition twisting motions across the central axis of every enzyme pair, Wakatsuki stated.
“This twist is nearly like a rachet that may push a completed product out or pull a brand new set of substances into the pocket the place the response takes place,” he stated. Collectively, the twisting and synchronization of the enzyme pairs enable them to repair carbon 100 occasions a second.
The ECR enzyme household additionally features a extra versatile department that may work together with many alternative sorts of biomolecules to provide a wide range of merchandise. However since they aren’t held collectively by molecular glue, they will’t coordinate their actions and due to this fact function rather more slowly.
“If we are able to enhance the speed of these subtle reactions to make new biomolecules,” Wakatsuki stated, “that might be a major bounce within the subject.”
From static photographs to fluid motion pictures
Up to now the experiments have produced static snapshots of the enzyme, the response substances and the ultimate merchandise in numerous configurations.
“Our dream experiment,” Wakatsuki stated, “can be to mix all of the substances as they move into the trail of the X-ray laser beam so we may watch the response happen in actual time.”
The crew truly tried that at SACLA, he stated, but it surely didn’t work. “The CO2 molecules are actually small, and so they transfer so quick that it’s exhausting to catch the second once they connect to the substrate,” he stated. “Plus the X-ray laser beam is so sturdy that we couldn’t hold the substances in it lengthy sufficient for the response to happen. Once we pressed exhausting to do that, we managed to interrupt the crystals.”
An upcoming high-energy improve to LCLS will possible remedy that downside, he added, with pulses that arrive rather more continuously – one million occasions per second – and may be individually adjusted to the perfect power for every pattern.
Wakatsuki stated his crew continues to collaborate with Erb’s group, and it’s working with the LCLS pattern supply group and with researchers on the SLAC-Stanford cryogenic electron microscopy (cryo-EM) services to discover a method to make this method work.
Researchers from the RIKEN Spring-8 Heart and Japan Synchrotron Radiation Analysis Institute additionally contributed to this work, which obtained main funding from the DOE Workplace of Science. A lot of the preliminary work for this research was carried out by SLAC summer season intern Yash Rao; interns Brandon Hayes, E. Han Dao and Manat Kaur additionally made key contributions. DOE’s Joint Genome Institute supplied the DNA used to provide the ECR samples. SSRL, LCLS, the Superior Photon Supply and the Joint Genome Institute are all DOE Workplace of Science consumer services.
Reference: “Intersubunit Coupling Allows Quick CO2-Fixation by Reductive Carboxylases” by Hasan DeMirci, Yashas Rao, Gabriele M. Stoffel, Bastian Vögeli, Kristina Schell, Aharon Gomez, Alexander Batyuk, Cornelius Gati, Raymond G. Sierra, Mark S. Hunter, E. Han Dao, Halil I. Ciftci, Brandon Hayes, Fredric Poitevin, Po-Nan Li, Manat Kaur, Kensuke Tono, David Adrian Saez, Samuel Deutsch, Yasuo Yoshikuni, Helmut Grubmüller, Tobias J. Erb, Esteban Vöhringer-Martinez and Soichi Wakatsuki, 25 April 2022, ACS Central Science.
DOI: 10.1021/acscentsci.2c00057
Post a Comment